Cytoscape transcription factor10/31/2022  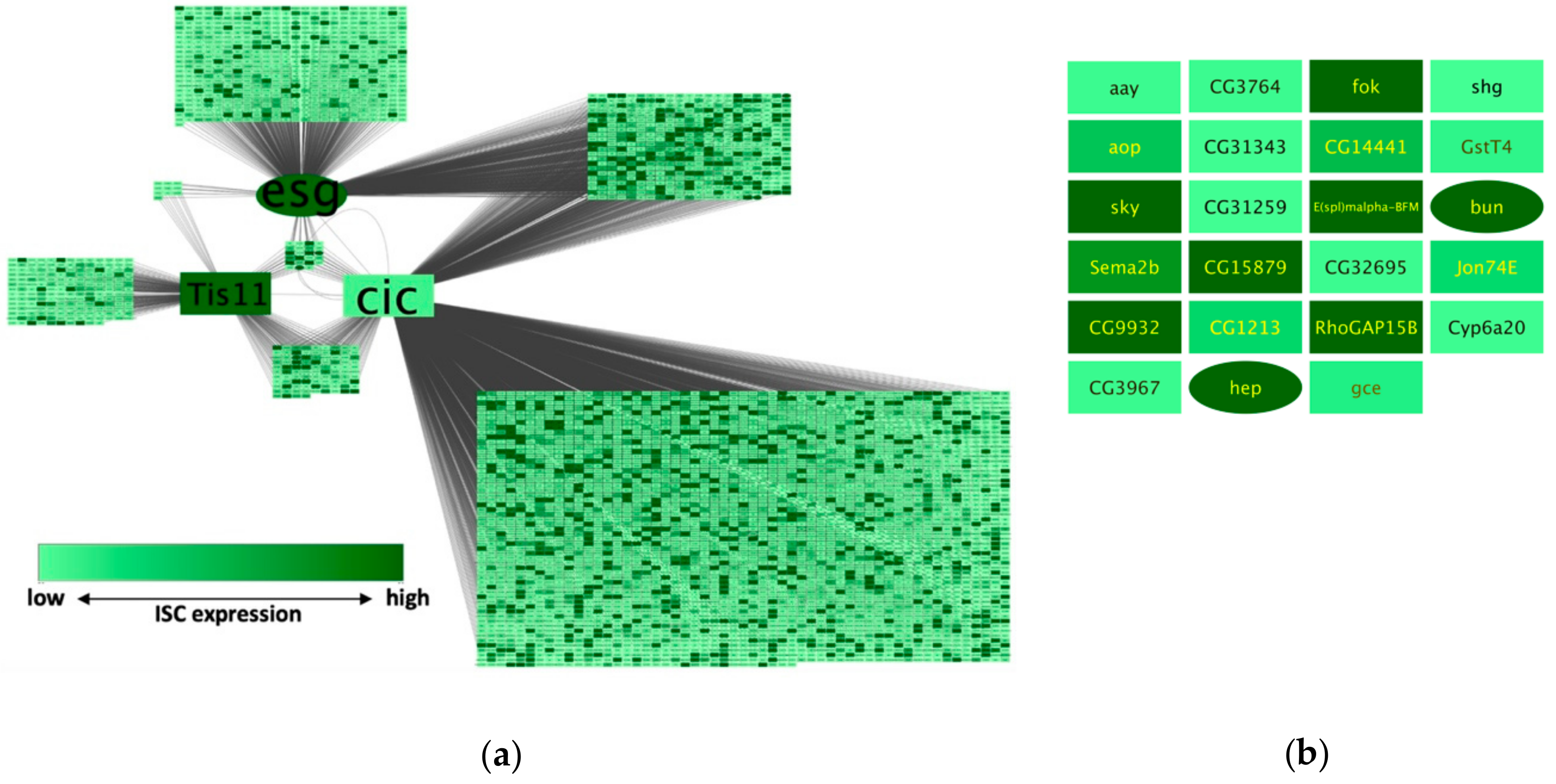
This will pop-up a node shape selection dialog.Double-click on the left node icon (a circle).This action will split the range of values with a slider down the middle with a node shape icon to either side of the slider.This action will pop-up a continuous shape selection dialog.This will create an empty icon in the Current Mapping row of the Shape section.Click the – select value – cell next to Mapping Type.This will produce a drop-down menu of available column names.Click the – select value – cell next to Column.cell next to the Shape row in the Style panel. We can use the significance values to change the shape of the nodes so that measurements we have confidence in appear as squares while potentially bad measurements appear as circles. We imported both expression measurement values and significance values for those measurements. Click OK to save the gradient adjustment dialog and verify that the nodes in the network reflect the new coloring scheme.This should produce a Blue-White-Yellow Color gradient like the image below, with min and max extremes colored black and green, respectively.You can also change the color of each handle by double-clicking or using the Node Fill Color selector button in the Handle Settings section.Finally, set the Maximum Color by double-clicking on the white, left-pointing triangle.Add a new handle by clicking Add, and drag that handle to 2.5.You can type the value in the Handle Position section to be more precise. Drag the white inverted triangle handle to approx 0.5.Double-click on the handle and set a color in the blue range. Drag the left-most, black inverted triangle handle along the top of the gradient.We’re going to build a basic blue-white-yellow gradient for our expression values.This will pop-up a gradient editing dialog. Click on the color gradient to change the colors.This action will produce a basic black to white color gradient.This will produce a drop-down menu of available mapping types.Click the – select value– cell in the Mapping Type section.Click the – select value – cell in the Column section.Click on the middle square ( Map.) next to the Fill Color row in the Styles panel.Under the Node Table, you should see your node listed with their expression values, as shown.ĭefine the node color of this visual style:.In the Node Table, You can limit the columns shown by going to the Show Columns button, and select the attributes gal1RGexp, gal4RGexp, and gal80Rexp by clicking on the checkbox.Select a node on the Cytoscape canvas by clicking on it.Now we will use the Node Table to browse through the expression data, as follows: Click the OK button to import the data.The default settings are usually correct, although you may need to change the Key column to indicate which column will be used to match with the network key column.Under the File menu, select Import → Table → File.In this case, there are three expression results per node. The remaining columns contain experimental data, two columns per experiment (one column represents the expression measurement and the second represents the significance value for that measurement), and one line per node.This column is optional, and the data is not currently used by Cytoscape, but including this column makes the format consistent with the output of many microarray analysis packages, and makes the file easier to read. The second column contains common locus names. 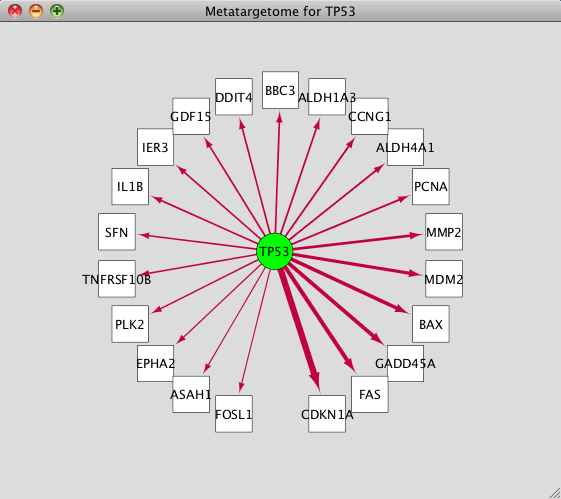
The first column contains node names, and must match the names of the nodes in your network exactly!.All columns are separated by a single comma character.You should note the following information about the file: 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
